The current subsections and their content are listed below:More.Sequence i The information is filed in different subsections. It also includes information pertinent to the sequence(s), including length and molecular weight. This section displays by default the canonical protein sequence and upon request all isoforms described in the entry. Superfamily database of structural and functional annotation More.SUPFAM i View protein in Pfam PF04348, LppC, 1 hit The PANTHER Classification System More.PANTHER i View protein in InterPro IPR007443, LpoA IPR008939, Lytic_TGlycosylase_superhlx_U IPR028082, Peripla_BP_I IPR011990, TPR-like_helical_dom_sf
Integrated resource of protein families, domains and functional sites More.InterPro i HAMAP database of protein families More.HAMAP i Gene3D Structural and Functional Annotation of Protein Families More.Gene3D i InParanoid: Eukaryotic Ortholog Groups More.InParanoid iĭatabase for complete collections of gene phylogenies More.PhylomeDB i The HOGENOM Database of Homologous Genes from Fully Sequenced Organisms More.HOGENOM i Curated Keywords - Domain i Signal Phylogenomic databasesĮvolutionary genealogy of genes: Non-supervised Orthologous Groups More.eggNOG i This subsection of the 'Family and domains' section provides information about the sequence similarity with other similarities iīelongs to the LpoA family. This subsection of the 'Family and Domains' section describes the position of regions of compositional bias within the protein and the particular type of amino acids that are over-represented within those bias i
#Pbp3 ecoli uniprot manual#
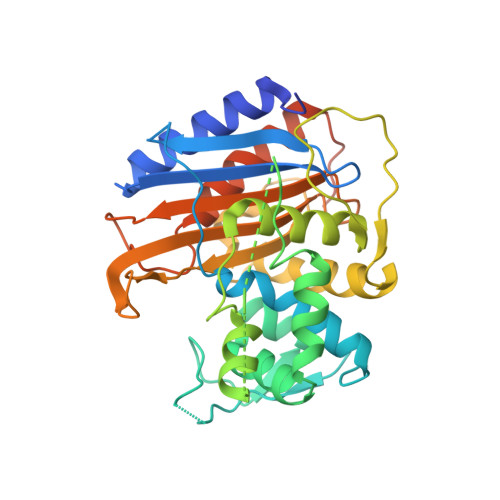
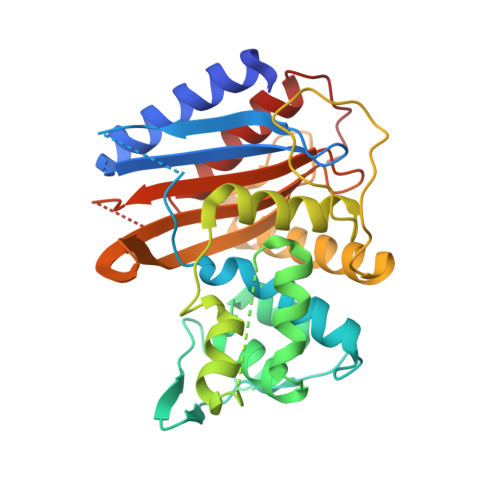
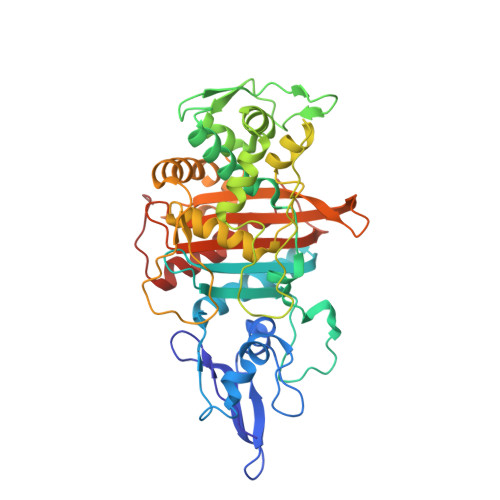
Information which has been generated by the UniProtKB automatic annotation system, without manual validation.Īutomatic assertion according to sequence analysis i This subsection of the 'Family and Domains' section describes a region of interest that cannot be described in other i This section provides information on sequence similarities with other proteins and the domain(s) present in a & Domains i Region Feature key Protein Data Bank in Europe - Knowledge Base More.PDBe-KB i SWISS-MODEL Repository - a database of annotated 3D protein structure models More.SMR iĭatabase of comparative protein structure models More.ModBase i This subsection of the 'Structure' section is used to indicate the positions of experimentally determined beta strands within the protein strand iīiological Magnetic Resonance Data Bank More.BMRB i Information inferred from a combination of experimental and computational evidence, without manual validation.Īutomatic assertion inferred from combination of experimental and computational evidence i This subsection of the 'Structure' section is used to indicate the positions of experimentally determined helical regions within the protein i Legend: Helix Turn Beta strand PDB Structure known for this area Show more details Hide details Feature key PRoteomics IDEntifications database More.PRIDE i PaxDb, a database of protein abundance averages across all three domains of life More.PaxDb i JPOST - Japan Proteome Standard Repository/Database More.jPOST i Keywords - PTM i Lipoprotein, Palmitate Proteomic databases S-diacylglycerol cysteine Sequence analysis This subsection of the PTM / Processing section specifies the position(s) and the type of covalently attached lipid group(s).More.Lipidation i
#Pbp3 ecoli uniprot Activator#
Penicillin-binding protein activator LpoA Add BLAST This subsection of the 'PTM / Processing' section describes the extent of a polypeptide chain in the mature protein following processing or proteolytic i PRO_0000169452 This subsection of the 'PTM / Processing' section denotes the presence of an N-terminal signal peptide i This section describes post-translational modifications (PTMs) and/or processing / Processing i Molecule processing Feature key It lists the nodes as they appear top-down in the taxonomic tree, with the more general grouping listed lineage iĬellular organisms › Bacteria › Proteobacteria › Gammaproteobacteria › Enterobacterales › Enterobacteriaceae › Escherichia › Escherichia coli This subsection of the Names and taxonomy section contains the taxonomic hierarchical classification lineage of the source organism. This is known as the 'taxonomic identifier' or 'taxid'.More.Taxonomic identifier i
This subsection of the Names and taxonomy section shows the unique identifier assigned by the NCBI to the source organism of the protein. This subsection of the Names and taxonomy section provides information on the name(s) of the organism that is the source of the protein i
